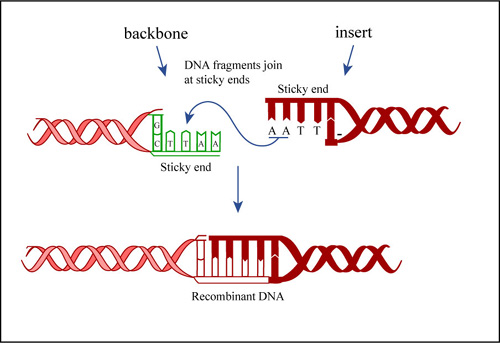
Insertion of DNA fragment to the vector is through the ligation process. Ligation is the joining of two fragment of DNA using the same endonuclease enzyme that cut both fragment of DNA.
The fragment which have been cut from vector and also gene of interest can be joint together by forming the phosphodiester boods between 5’-phosphoryl of DNA terminus and 3’-hydroxyl end of another. A cofactor also involved such as ATP and NAD+ in this reaction. Sometimes, the vector may self-ligation which recircularized the vector DNA so this problem can be overcome by using alkaline phosphatase which remove the 5’-phosphate group. Self-ligation can overcome as the end of DNA strand doesn’t exist 5’-phosphoryl end to ligate.
The fragment which have been cut from vector and also gene of interest can be joint together by forming the phosphodiester boods between 5’-phosphoryl of DNA terminus and 3’-hydroxyl end of another. A cofactor also involved such as ATP and NAD+ in this reaction. Sometimes, the vector may self-ligation which recircularized the vector DNA so this problem can be overcome by using alkaline phosphatase which remove the 5’-phosphate group. Self-ligation can overcome as the end of DNA strand doesn’t exist 5’-phosphoryl end to ligate.
http://ocw.mit.edu/courses/biological-engineering/20-109-laboratory-fundamentals-in-biological-engineering-fall-2007/labs/mod1_3/
Type of ligases
a) T4 DNA ligase
This DNA ligase is bacteriophage T4 which is single polypeptide that ligate cohesive or “sticky” ends and blunt end of DNA. It needed ATP as energy source and maximal pH range is 7.5-8.0. Mg+ ion is also required and the optimal concentration is 10mM. Joining blunt end is more complex and significant slower.
b) E.coli DNA ligase
This ligase ligate cohesive and “sticky” DNA and cannot ligate blunt-end DNA unless polyethylene glycol or Ficoll This enzyme acquire energy by cleaving nicotinamide adenine dinucleotide (NAD) to create phosphodiester bond.
c) Thermostable DNA ligase
Ligase that come from thermophilic bacteria that catalyses NAD-dependant ligation in duplex DNA structures. This can be added to PCR reaction.
Linker
|
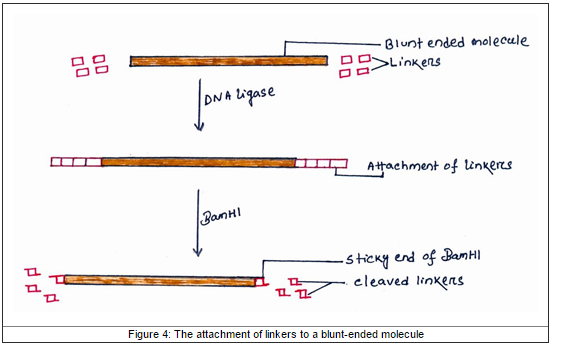
It is short synthetic palindromic sequences that ligated to the other DNA molecules that used to introduce restriction sites in the DNA after ligation. Linker will attach to the ends of blunt-end of DNA by ligation with DNA ligase. Then the endonucleases enzyme will digest the linker so that it can insert into the vector that contain the complementary ends. For example, EcoRI cleaves the chains at the recognition sequences, produce many linker that have been cleaved by EcoR1 and original fragment of DNA, the cleaved linker is attached to the original DNA fragment by ligation. So, the modified fragment can ligated into vector that also cleaved by EcoR1.
|
Adaptor
|
http://www.discoverbiotech.com/wiki/-/wiki/Main/DNA+Ligation+and+Transformation |
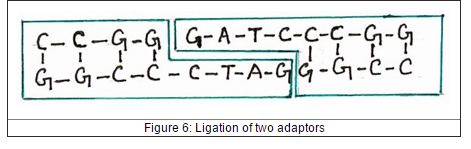
Adaptor also same with linker are chemically synthesized oligonucleotide which ligate to the ends of DNA chains. But, adaptor is not same with linker that adaptors has one sticky end already. The blunt end of adaptor will ligates to the blunt end of fragment of DNA which needed insert in vector to produce new DNA molecule with sticky end. But, the individual adaptor can base pair with each other to form new dimers.
|
|
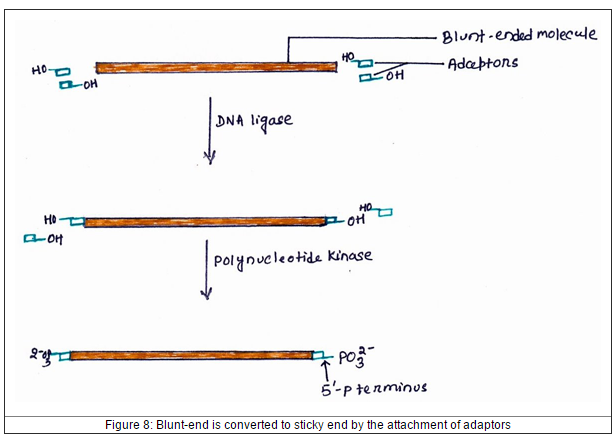
To prevent this problem, adaptor molecules are created so that blunt end ligates with DNA fragment is same as natural DNA while sticky end is different. The 3’-hydroxyl terminal unchanged but the 5’-phosphoryl terminal is changed to 5’-hydroxyl group. Then, the enzyme, polynucletide kinase which has been used to catalyze reaction of transfer the phosphate group from ATP to 5'-hydroxyl terminal. Now, the adaptor can attached to both DNA fragment and vector which cut with the same restriction enzyme. |
http://www.discoverbiotech.com/wiki/-/wiki/Main/DNA+Ligation+and+Transformation
|